Import external Datasets to the database
Datasets that have been obtained outside of the Visiopharm platform can be imported into the database. This allows for the data to be utilized in the various analytical tools available in the Visiopharm platform, such as the Explore/QC tool.
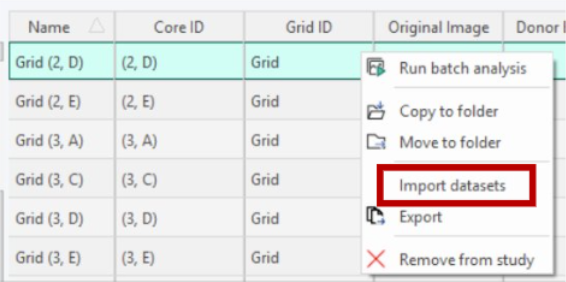
To import a dataset, right-click on the same image where the data was originally aquired.
Supported Formats
Two formats are supported:
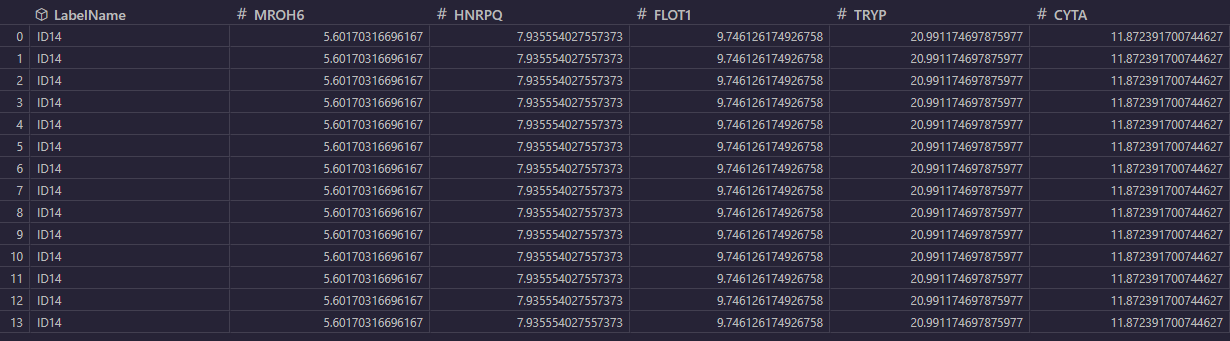
- Perseus format — Expects a Perseus-style TSV where the first data row contains column type markers (#!{Type}E for expression/numeric columns, T for text columns). A Protein.Group text column is required. During import, the importer matches E-type column names to ROI region-type names in the image's overlay. Each ROI object found in the image gets its corresponding column of values mapped into the result. Rows in the output correspond one-to-one with the data rows (excluding the type-marker row), and columns correspond to matched ROI objects. Unmatched regions are reported to the user.
An example of how the file should look like:
- Generic TSV — For plain tab-separated files (.txt, .tsv). The file is read as-is (tab delimiter, with header row) into a DataList and re-saved in the internal CSV format. No column/row remapping is performed.
Supported file formats for datasets are .txt, .csv, and .tsv.
Row Alignment with Per-Object Results
Per-object results, which is sourced from Object Info, produce one row per detected object, where each row carries identifying metadata (label type/name, ROI type/name, bounding-box envelope, object ID). When importing external TSV data to complement these results, the imported rows must align by position: row N in the imported dataset corresponds to object N in the per-object result list.
There is no key-based join: the Perseus importer achieves alignment by iterating over ROI objects in the overlay in the same order they appear, and writing one column of values per object. For the generic importer, make sure that the row order in the TSV matches the object enumeration order produced by the analysis.
Be aware that the name of the column will be named after the dataset filename, as can be seen in the images below: