t-SNE plots
A t-SNE (t-distributed Stochastic Neighbor Embedding) plot will show the combination of high-dimensional data (that is, with more than two output variables) in a computed two-dimensional map. t-SNE uses the original distances in the high-dimensional space to calculate the t-SNE coordinates. In simple terms, similar data points in the high-dimensional space tend to group in the t-SNE coordinates as the weight of the distances is preserved, but not the actual distance. When working with t-SNE plots, use Color by as the first step, then interact manually with the data points, and finally tune the hyperparameters.
Creating t-SNE plots
After selecting the variables to visulize with a t-SNE plot, there are four t-SNE parameters that can be adjusted. Three hyperparameters; The perplexity, Learning rate and Max iterations, as well as a Panning Only option, which is relevant for the subsequent inspection of the results and can be toggled on or off.
Perplexity can be interpreted as a smooth measure of the effective number of neighbors, where the larger the perplexity, the more non-local information will be retained in the dimensionality reduction result.
Learning rate is a factor contributing the gradient update, which includes an exponentially decaying sum of previous gradients as a way of avoiding poor local minima. A very large learning rate will demand more iterations to ensure convergence.
Max iterations denotes how many iterations will be done. If the number of iterations is set very high and the plot has reached convergence, plotting can be stopped early by pressing Stop.
After selecting the desired variables under Select Variables, an initial t-SNE plot can be generated using the default parameters by clicking the button ![]() .
Once t-SNE is running, the plot will update live in the plot window as the iterations progress.
.
Once t-SNE is running, the plot will update live in the plot window as the iterations progress.
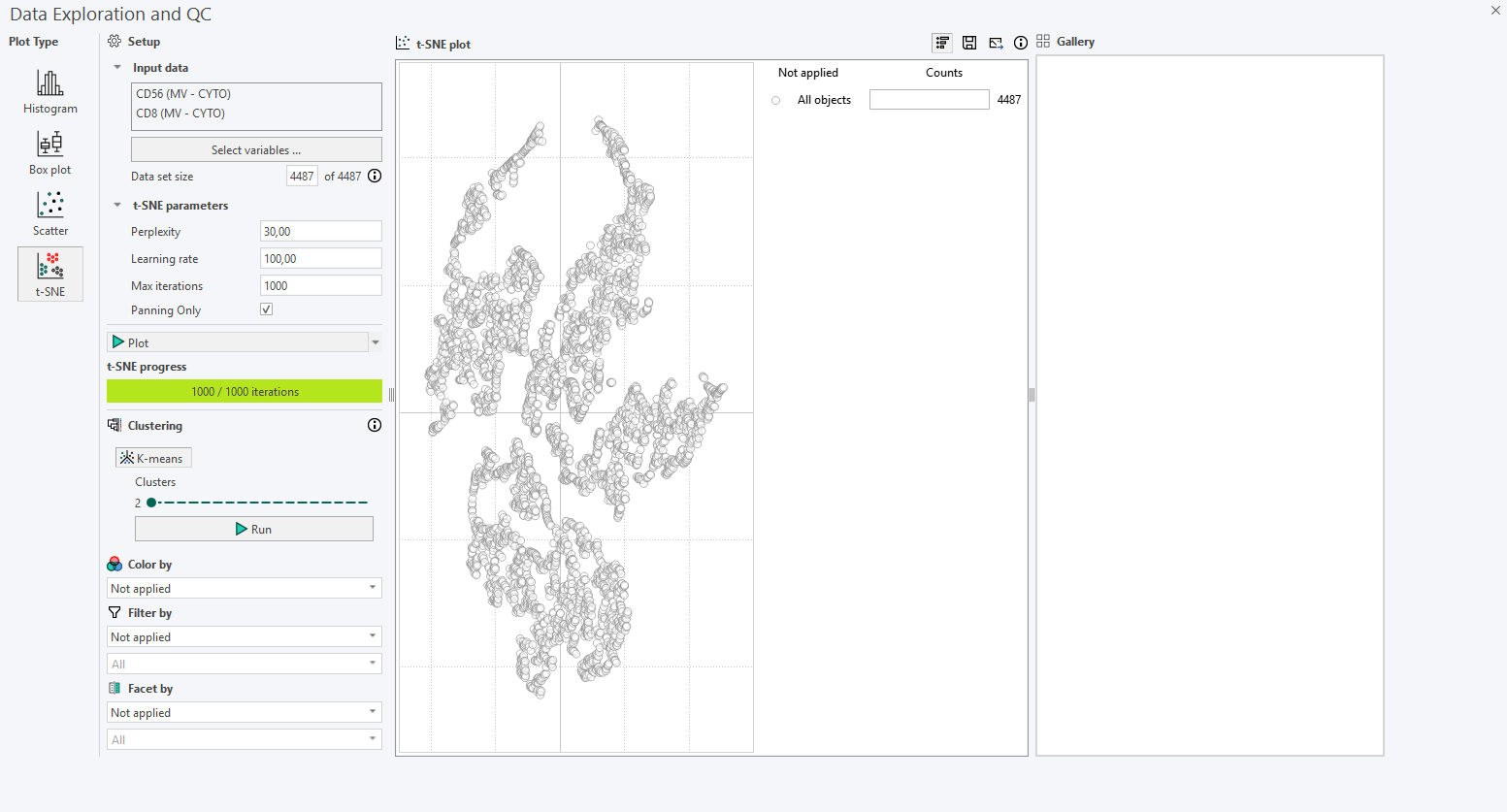
After running the first t-SNE plot with default parameters, inspect the results:
- Look for any shapes that are forming subpopulations.
- Use Color by with continuous or categorical variables to reveal potential subpopulations. When applicable, the Min/Max percentile can also be adjusted. If a parent category is selected to color the points (e.g., biomarker intensity), a secondary dropdown menu becomes available for selecting the specific biomarkers to apply thresholding to, as shown below.
- Use Filter by with categorical variables to investigate a certain sub-selection(s) of the full data set and look if any variable can explain the plot.
- Use the interaction with the points to move the image viewer to that point.
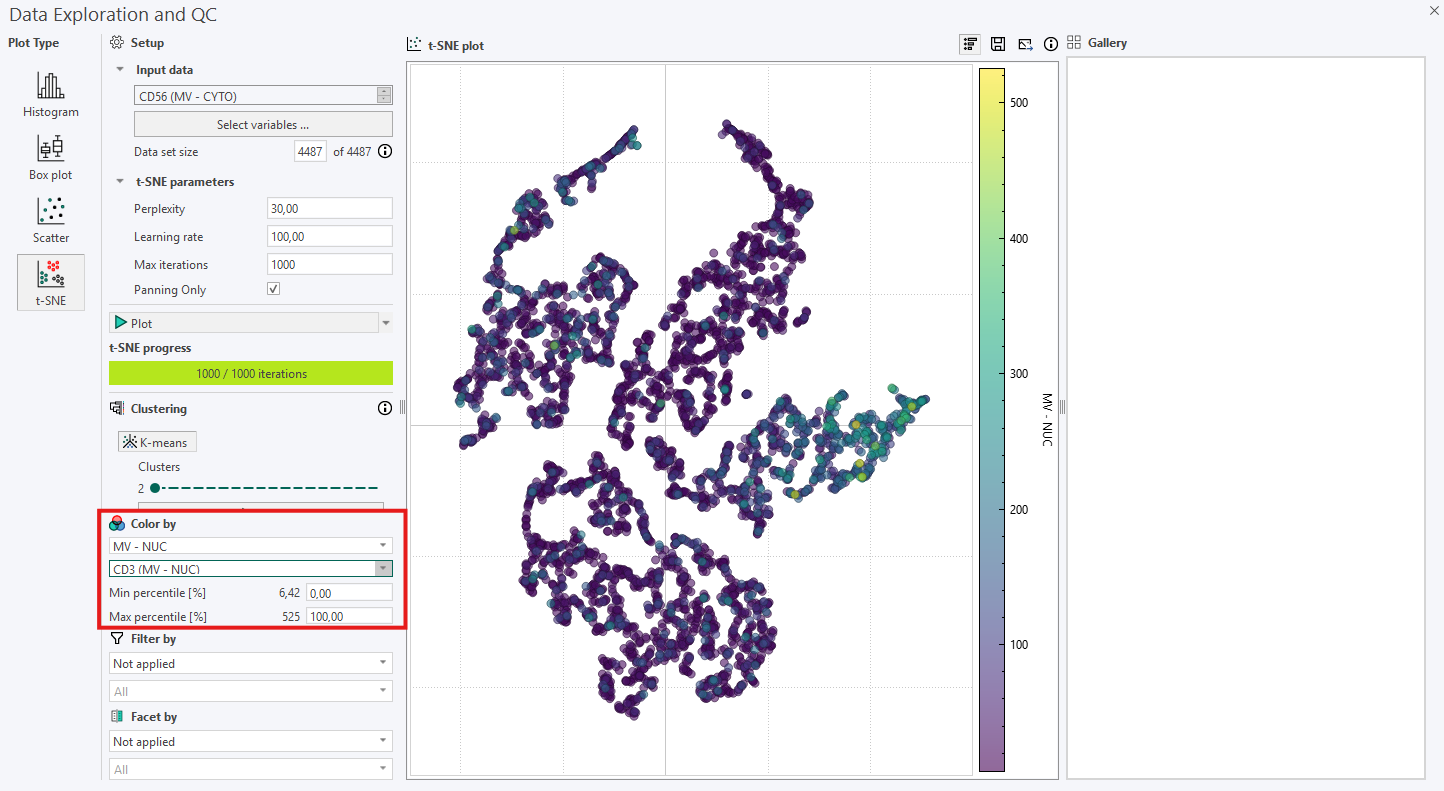
If the first inspection does not reveal any relevant information, remember that the hyperparameters will affect the plot. Try to re-run the algorithm while adjusting the hyperparameters systematically in the following order:
- Perplexity: If no clear subpopulations have formed, try to adjust the perplexity. This parameter controls how the algorithm balances attention to the local and global aspects of the original high-dimensional data.
- Iterations: Different data sets can require different numbers of iterations to converge. Increase the number of iterations if the t-SNE plot looks “stretched” or “folded” onto itself. Increase until cluster stability is reached, e.g., 1000 to 5000 where defined clusters are separated by equal spaces.
- Learning rate: This parameter should be adjusted last, and it controls how large steps/adjustments the algorithm is allowed to make at each iteration.
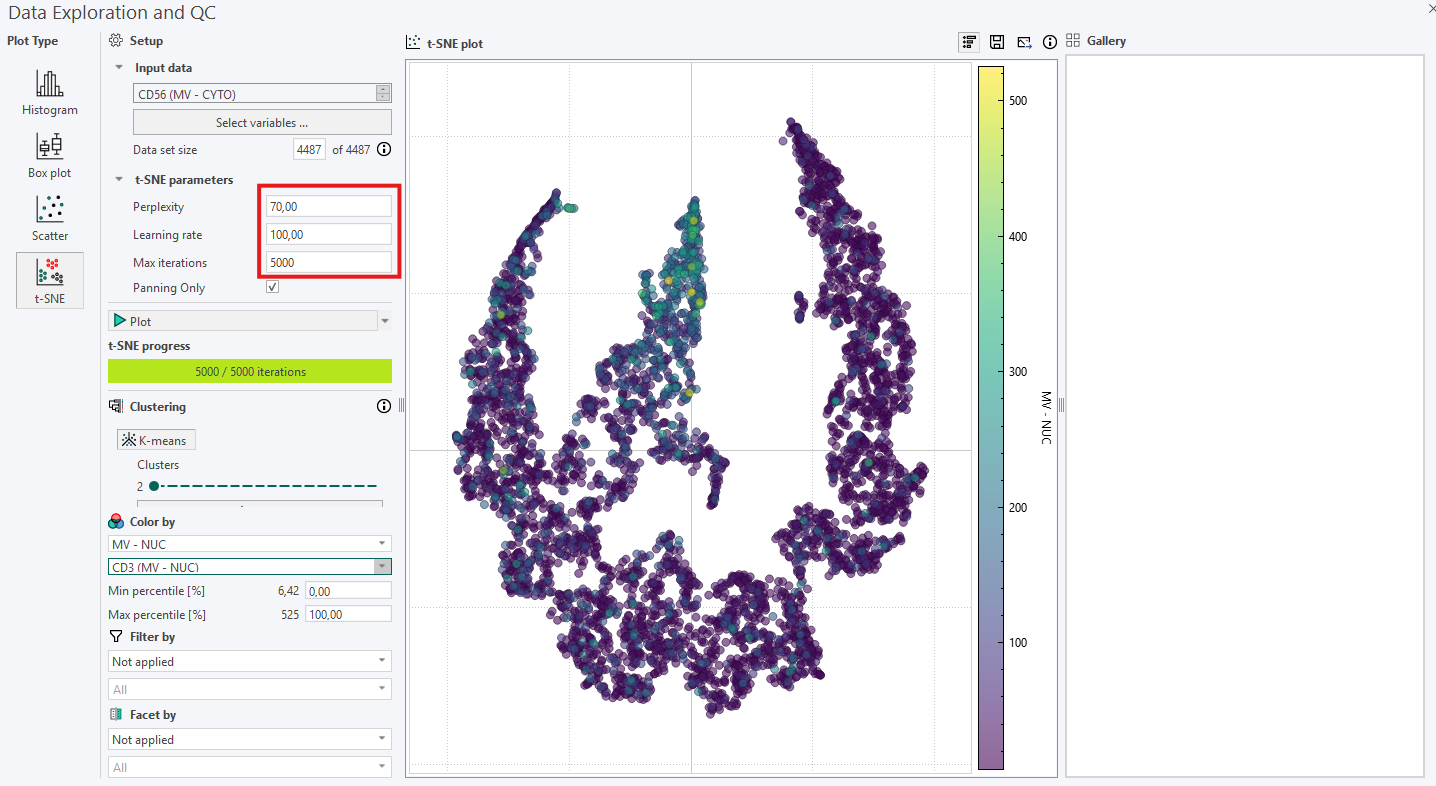
Outliers in t-SNE plots are points that do not belong to the group where t-SNE placed them. To spot outliers, start by coloring the data one by one with the same output variables used as input to the t-SNE algorithm. The goal is to find a variable that will give clustered points the same color. Outliers can occur as points with a distinct color inside an otherwise homogeneously colored collection of data points but remember to manually verify this by interacting with the data point and looking at the data in the image viewer.
k-means clustering can be used to identify and group similar data points in a lower-dimensional space. This is done by asssigning each point to a cluster, to which it is most similar to. Increasing the number of clusters, k, will allow for identification of less obvious patterns in the data, but might also lead to non-descriptive clusters. Values for k can be between 2 and 20.
Facet by can be used to divide the t-SNE plot into subplots based on Label, ROI, Image Name, or Folder Name. In this example, the t‑SNE plot is faceted by ROI, resulting in a Stroma and a Tumor subplot.
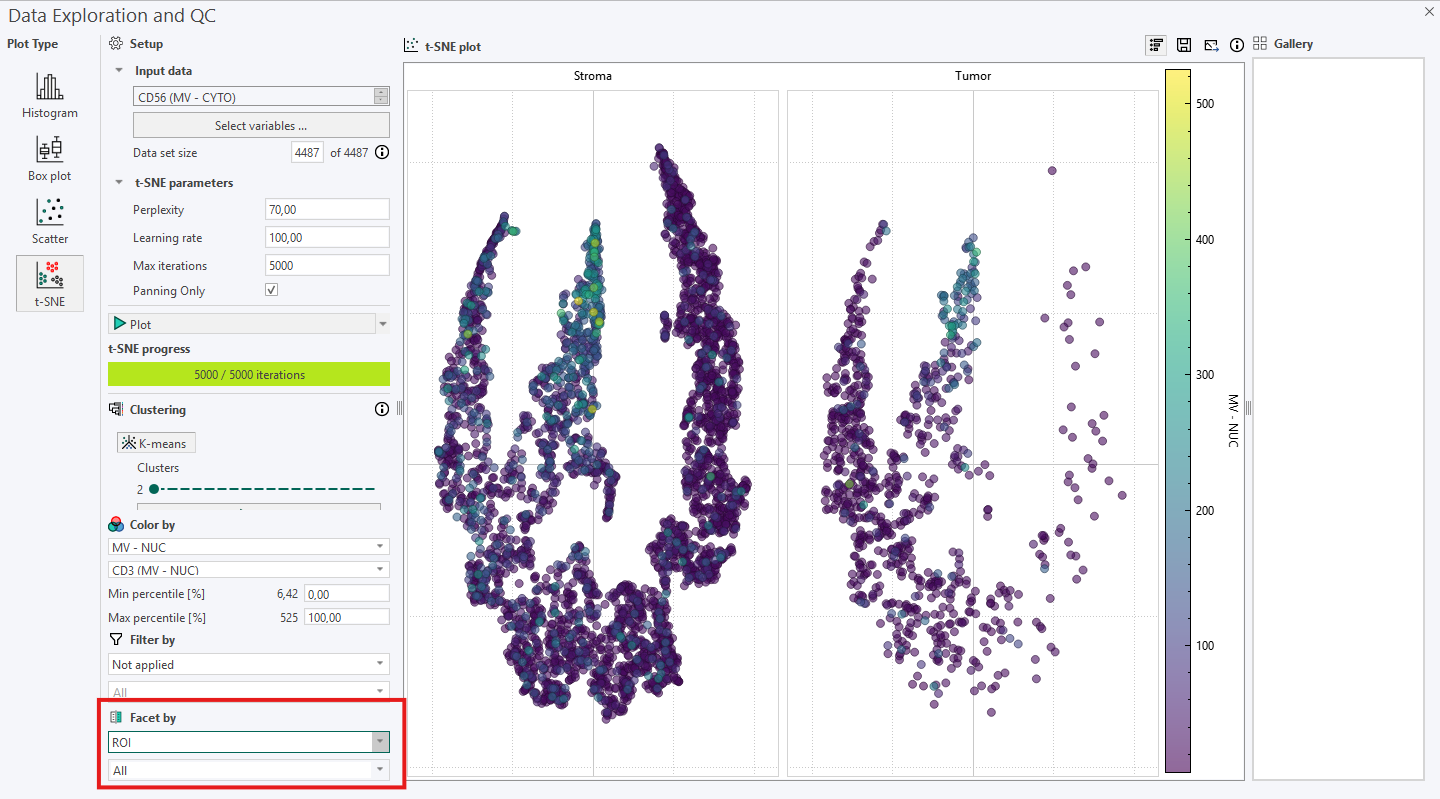
When using t-SNE remember the following:
- t-SNE subpopulation sizes carry no meaning.
- t-SNE distances are not representative of the original data differences.
- Random noise in the original data is not always random in t-SNE coordinates.
- Shapes only sometimes appear, so use color by to find structures.
- Run several experiments varying (at least) the perplexity.
Saving the plots can be done either by clicking ![]() to save the current plot data to the database, or by clicking
to save the current plot data to the database, or by clicking ![]() to save an image of the t-SNE plot from the current view to a specified folder.
to save an image of the t-SNE plot from the current view to a specified folder.
Inspecting t-SNE Results
Legend bar
In versions 2024.07 and newer, an interactive legend appears to the right of the t-SNE plot. Initially, the legend displays the total number of cells in the plot. For example, the following plot contains 4487 cells.
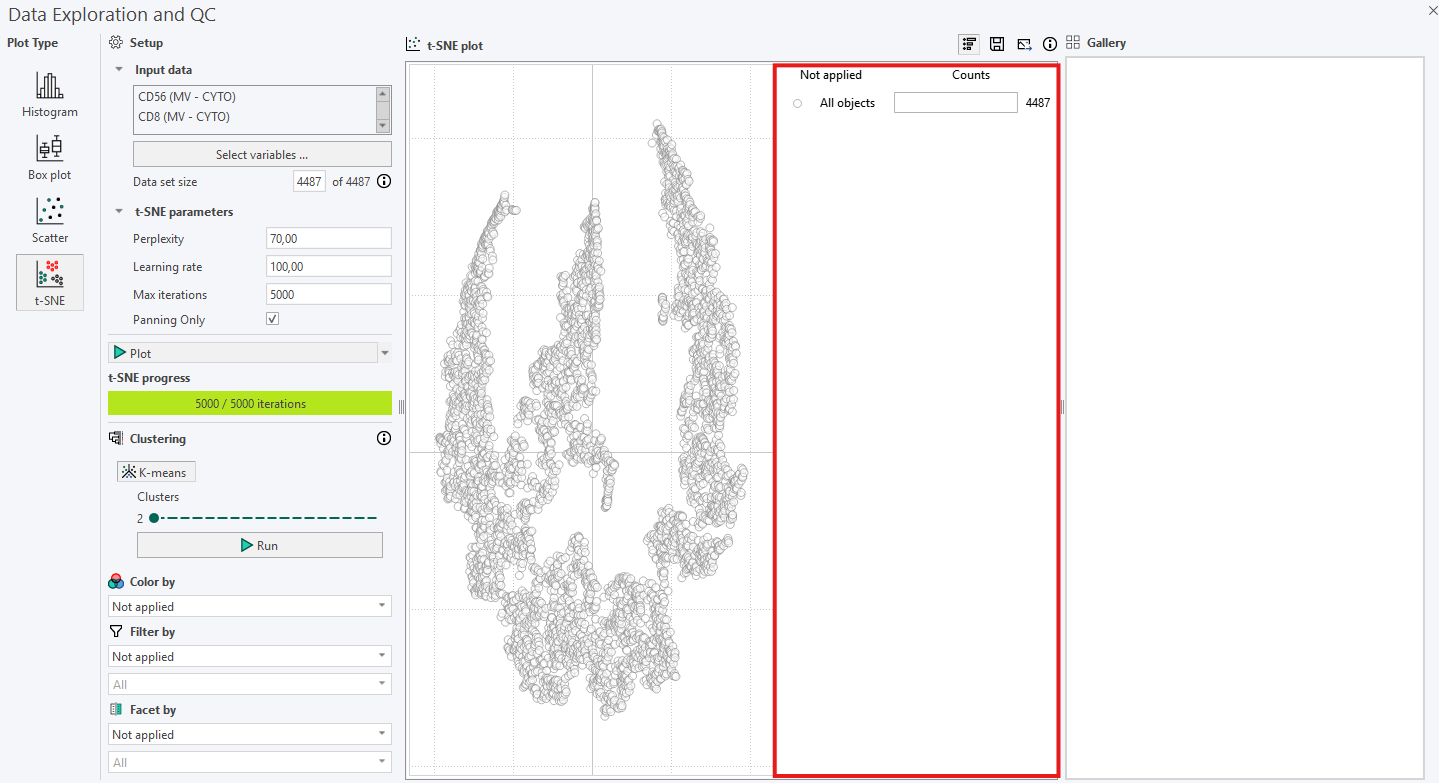
Cells can be colored according to the Region of Interest (ROI) they originate from using Color by. When this option is enabled, the legend automatically updates to show the number of cells in each ROI for the selected sample image. The same principle applies when coloring cells by Labels.
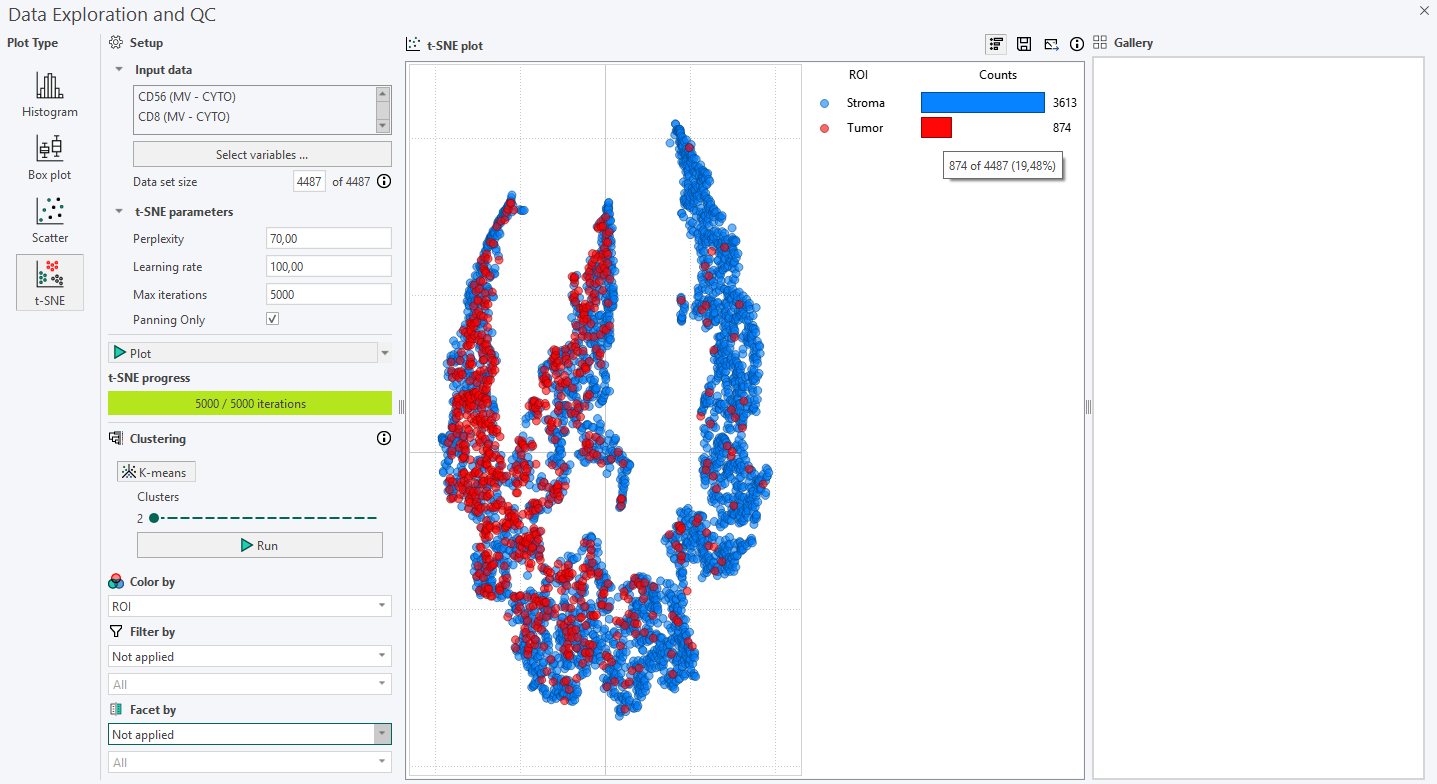
The legend bar can be shown or hidden by right-clicking directly on it. Right-clicking again in the same area will make it reappear.
If only the ROI names are needed, without the cell counts, the count bars can be collapsed and restored at any time by clicking on ![]() .
.
Hovering over a category in the legend bar will display the number of cells in that specific ROI out of the total number of cells, along with its percentage of the total cell count.
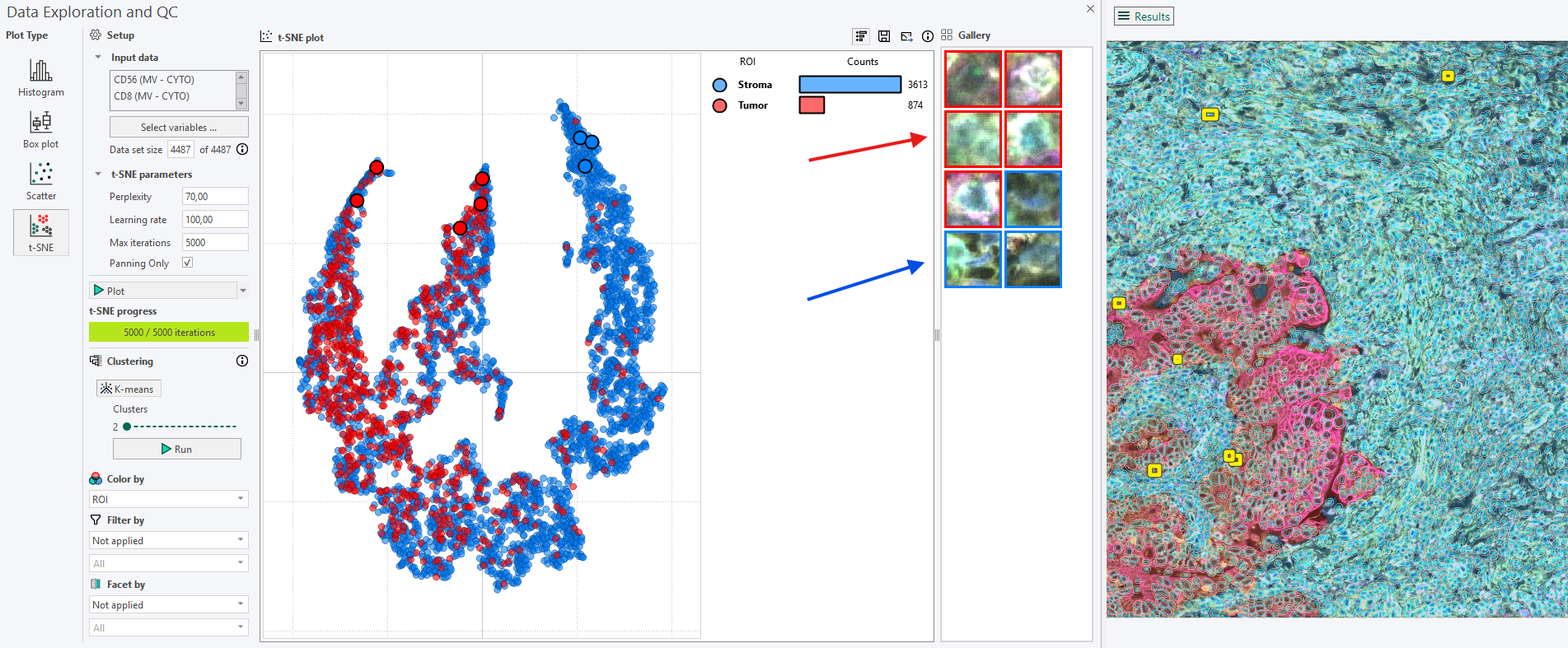
The Gallery
The Gallery is located to the right of the legend bar and is used to display how the cells highlighted in the t-SNE plot appear in the corresponding image. Cells can be added to the Gallery in several ways.
1. Extracting a random sample from the legend bar
From the legend bar, clicking on an ROI label (e.g., the circular ROI indicator “Tumor”) extracts a random sample belonging to that ROI.
The selected cells are shown as yellow squares in the image and are simultaneously highlighted in the t-SNE plot.
Clicking the label again or pressing the N key generates a new random sample.
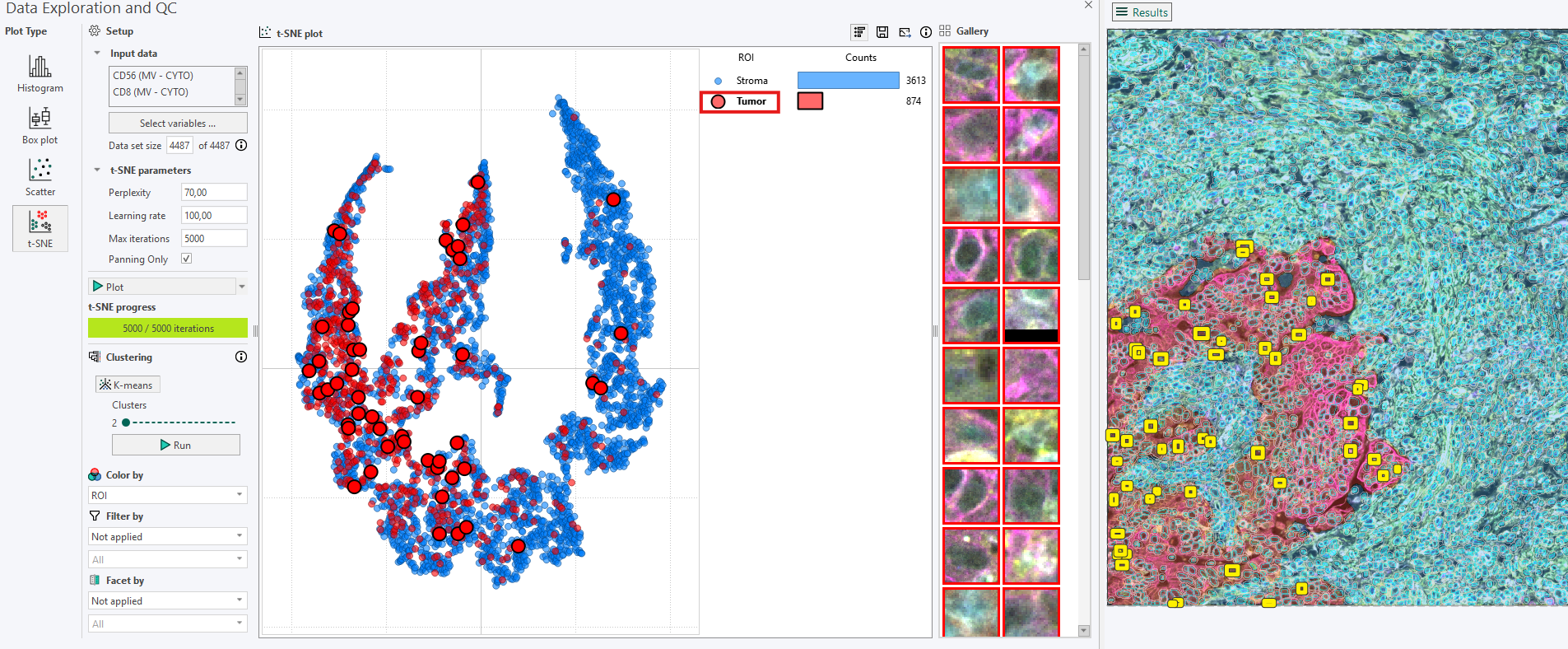
2. Highlighting all cells from a category
Clicking on the count bar in the legend highlights all cells belonging to the selected ROI in the image, while still highlighting a random sample in the t-SNE plot and the Gallery. Be aware that, for very large datasets, this can become visually dense, which is why the random-sample option is often preferable.
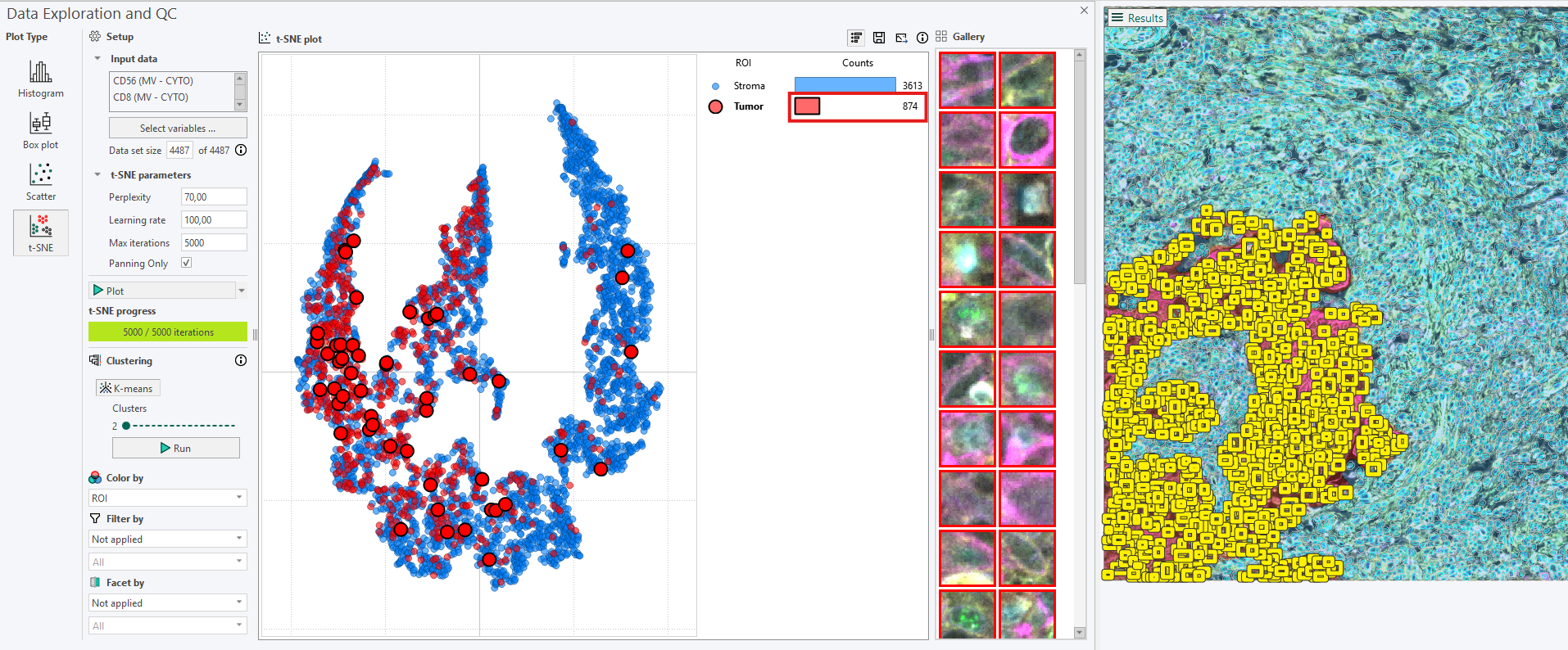
3. Viewing selected points of interest
Clicking on a point in the t-SNE plot or in the Gallery highlights the corresponding cell in the image, and vice versa.
Using Ctrl + Click makes it possible to select multiple points of interest in the t‑SNE plot, all of which will be displayed and highlighted in both the Gallery and the image.
The color surrounding each image in the Gallery indicates the category (e.g., ROI) to which the cell belongs.
Navigation through the Gallery image crops can be performed using the left/right arrow keys.
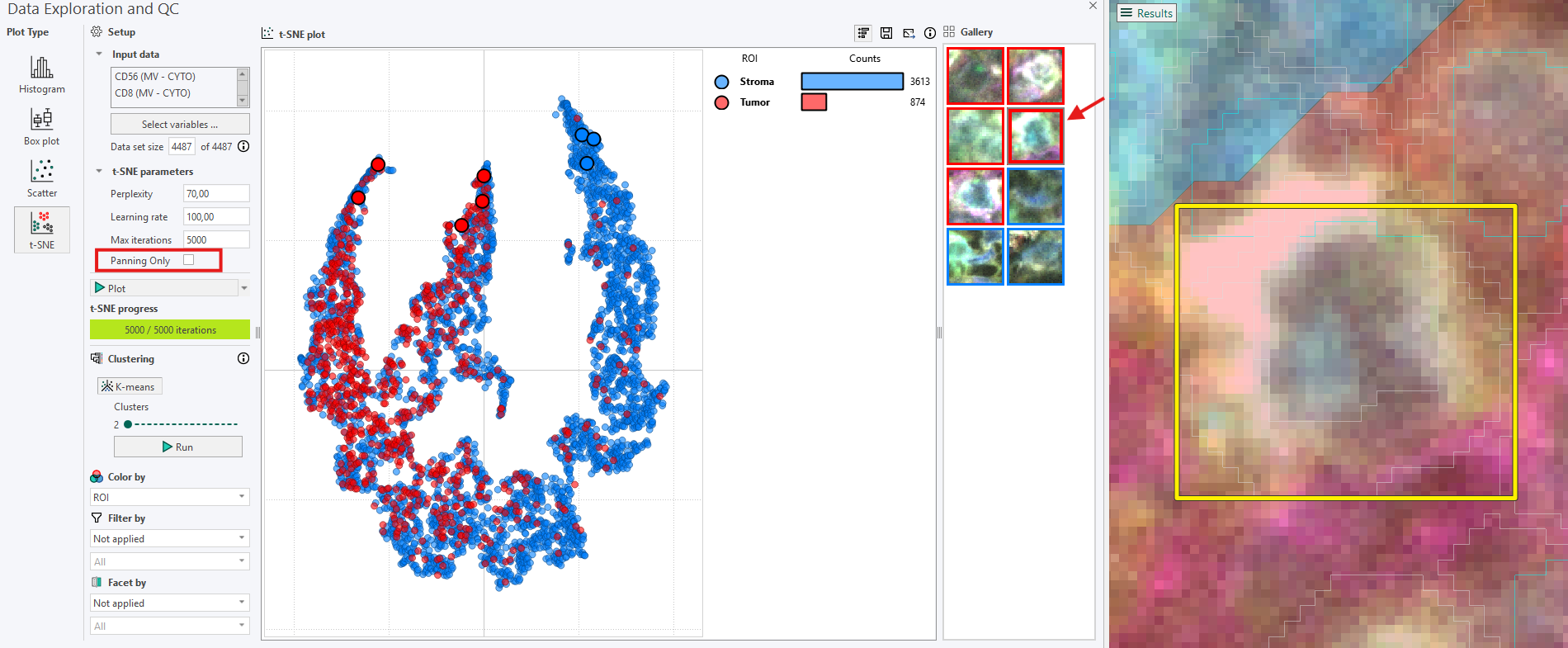
Panning Only mode
The Panning Only option, found in the t-SNE parameters section, controls how zooming behaves when interacting with the Gallery.The default setting is to have Panning Only enabled, but it can easily be toggled on or off as needed.
When Panning Only is off: Clicking on a Gallery image crop zooms directly into that cell. To inspect spatial context, the view must be zoomed out again.
When Panning Only is on: The zoom level chosen by the user is preserved. Clicking on a Gallery image crop moves the view so that the crop is centered, but does not zoom directly into the cell.
For more information please refer to: the original paper on t-SNE or How to Use t-SNE Effectively.